SCIENTIFIC ACHIEVEMENT
At the Advanced Light Source (ALS), researchers clarified the atomic-level mechanism that enables bacteria to switch light harvesting off and on in response to potentially damaging overexposure to light.
SIGNIFICANCE AND IMPACT
The results could have long-range implications for artificial photosynthesis and optogenetics—the use of light to selectively activate biological processes.
A prototype of photosynthesis
Cyanobacteria are water-dwelling microbes capable of absorbing sunlight and converting it into chemical energy through photosynthesis. Long ago, ancient versions of these bacteria were incorporated into plant cells, where they eventually evolved into chloroplasts, the organelles responsible for carrying out photosynthesis in green plants. Today, in seeking to develop artificial photosynthesis to harness the sun’s abundant energy, scientists look to cyanobacteria to better understand the nuts and bolts of how natural photosynthesis works.
Cyanobacterial “off switch”
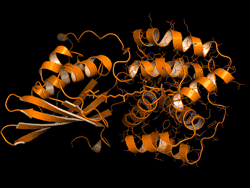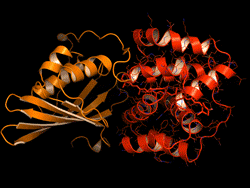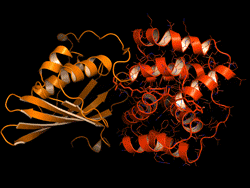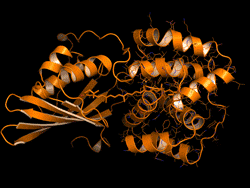
One topic of interest is how cyanobacteria respond to too much light. If a sunlight-harvesting system becomes overloaded with absorbed solar energy, it most likely will suffer some form of damage. Nature has solved the problem in cyanobacteria through a protective mechanism—an energy-quenching “off switch” in which excess solar energy is safely dissipated as heat.
In previous work on this topic, researchers focused on orange carotenoid protein (OCP), a light-sensitive protein embedded in cyanobacterial membranes. OCP was found to be triggered by the shifting of a hydrocarbon chain—a carotenoid molecule—between the two main OCP domains—the N-terminal domain (NTD) and the C-terminal domain (CTD). As a result of this translocation, OCP shifts from an “orange” light-absorbing state (OCPO) to a “red” photoprotective state (OCPR).
Fluorescence recovery protein
In this work, the researchers studied another protein complex, called fluorescence recovery protein (FRP), to see what role it plays in switching photosynthesis back on. The photoprotective OCPR state is metastable; it relaxes back to OCPO slowly in darkness, but instantaneously in the presence of FRP. Thus, determining the exact structural changes that accompany OCPR formation and its interactions with FRP is critical for a complete mechanistic understanding of the regulation of photosynthesis in cyanobacteria. But although the structure of free FRP has been solved, its detailed interactions with OCP and its precise mechanism of photoactivation has not been well understood.
Following the footprints
To discover how FRP facilitates the resumption of photosynthesis, the researchers performed x-ray footprinting experiments at ALS Beamline 5.3.1. The technique involves exposing FRP–OCP complexes to x-rays while in solution. The high-flux-density x-ray beams ionize water molecules in the solution, generating hydroxyl radicals that label all solvent-accessible areas of the proteins. Timing the x-ray pulses to hit at various intervals after initial excitation of the complexes with blue light provides time-resolved data. The hydroxyl-labeled protein complexes were then fragmented by proteolytic digestion and analyzed by mass spectrometry at the Joint BioEnergy Institute, to produce site-specific maps of the FRP–OCP interaction areas inaccessible to the solvent (the interaction “footprint”) at a single-residue resolution.
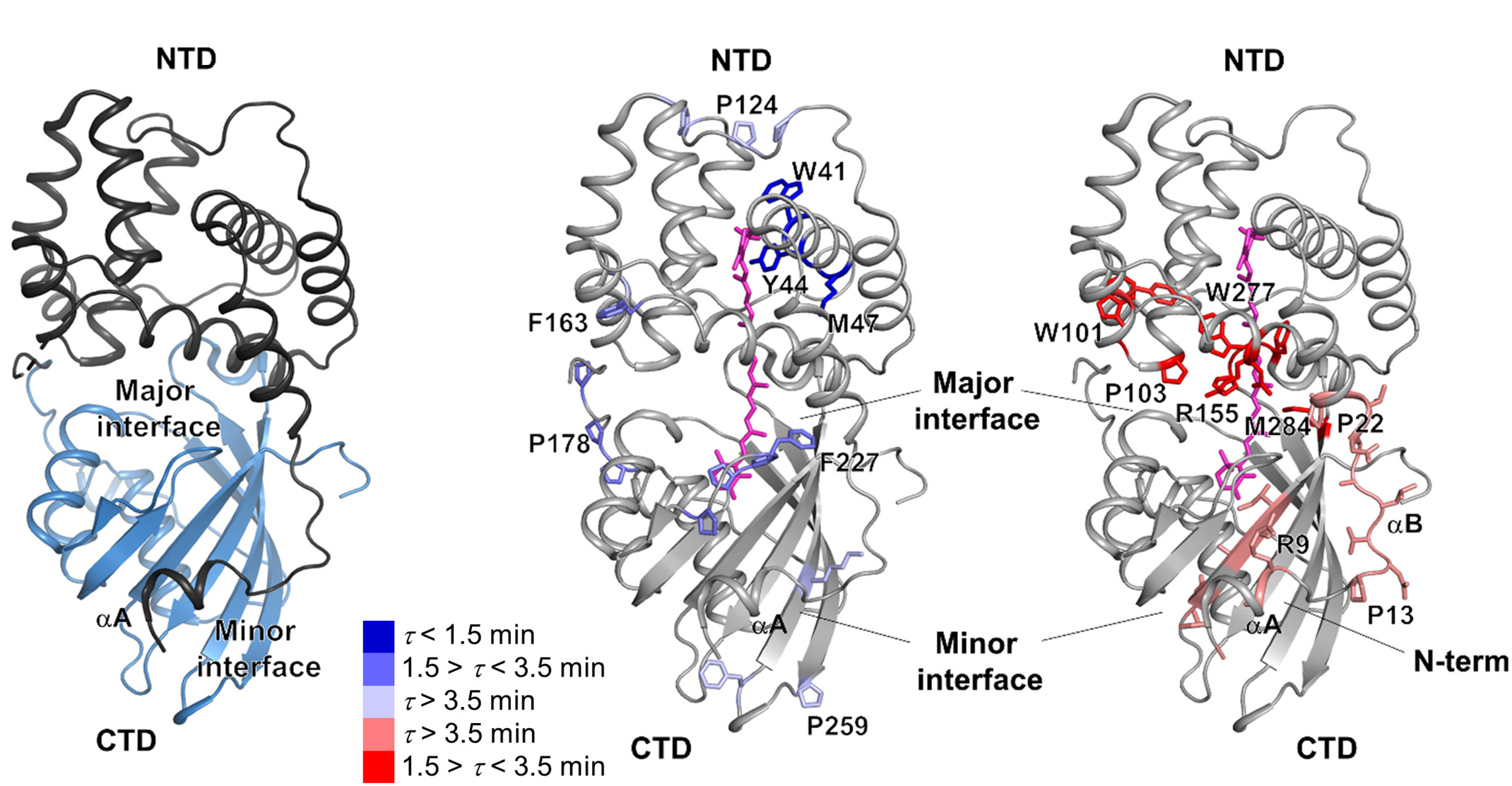
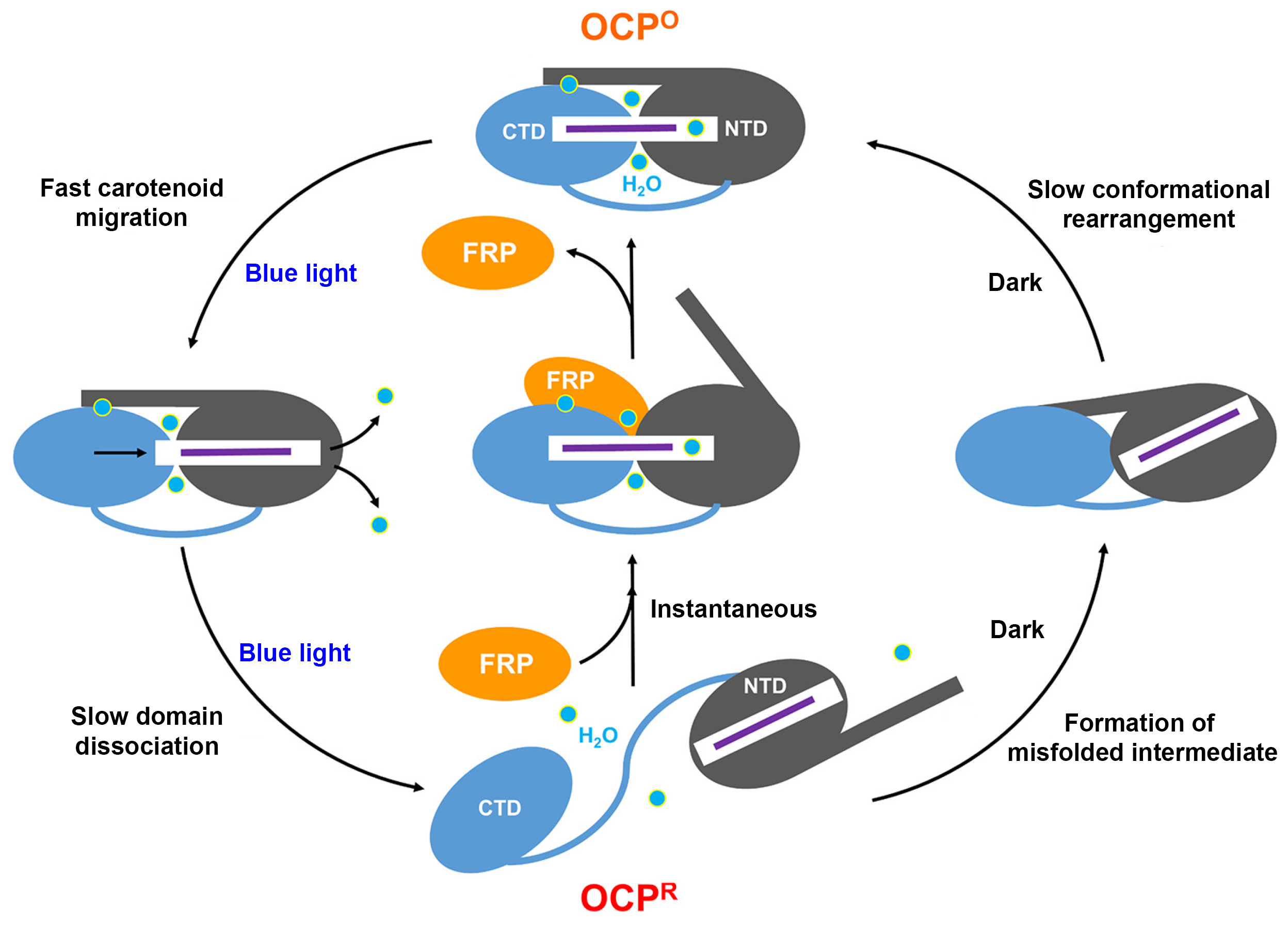
A working model of photoprotection
Based on the results, the researchers were able to develop a working model of how FRP creates a “shortcut” to OCP relaxation by holding the NTD and CTD together like a scaffolding. They also found that, without FRP, this particular OCP variant forms a misfolded intermediate that may slow down the relaxation process. In general, understanding the mechanisms controlling the thermal dissipation of excess light energy has long-range implications in synthetic biology, for example in the design of selective, light-activated switches for biological processes (optogenetics), or for finer control of photoprotection and consequent enhancement of photosynthetic efficiency.
Contact: Sayan Gupta
Researchers: S. Gupta, S.G. Remesh, X.A. Feng, L.-J.G. Chan, C.J. Petzold, and C.Y. Ralston (Berkeley Lab); M. Sutter and CA. Kerfeld (Michigan State University and Berkeley Lab); and M.A. Dominguez-Martin and H. Bao (Michigan State University).
Funding: National Science Foundation, National Institutes of Health, and the U.S. Department of Energy (DOE), Office of Science, Office of Biological and Environmental Research. Operation of the ALS is supported by the DOE Office of Science, Basic Energy Sciences Program.
Publication: S. Gupta, M. Sutter, S.G. Remesh, M.A. Dominguez-Martin, H. Bao, X.A. Feng, L.-J.G. Chan, C.J. Petzold, CA. Kerfeld, and C.Y. Ralston, “X-ray radiolytic labeling reveals the molecular basis of orange carotenoid protein photoprotection and its interactions with fluorescence recovery protein,” J. Biol. Chem. 294, 8848 (2019), doi:10.1074/jbc.RA119.007592.
See also the Berkeley Lab Biosciences highlight, “X-ray Footprinting Reveals Molecular Basis of Orange Carotenoid Protein Photoprotection.”
ALS SCIENCE HIGHLIGHT #400