SCIENTIFIC ACHIEVEMENT
Researchers used soft x-ray tomography at the Advanced Light Source (ALS) to gain a 3D whole-cell view of how insulin-producing pancreatic cells react upon exposure to glucose and a diabetes drug.
SIGNIFICANCE AND IMPACT
The approach enables direct quantification of intracellular responses before, during, and after cell stimulation, providing new insights into how drugs alter cell function.
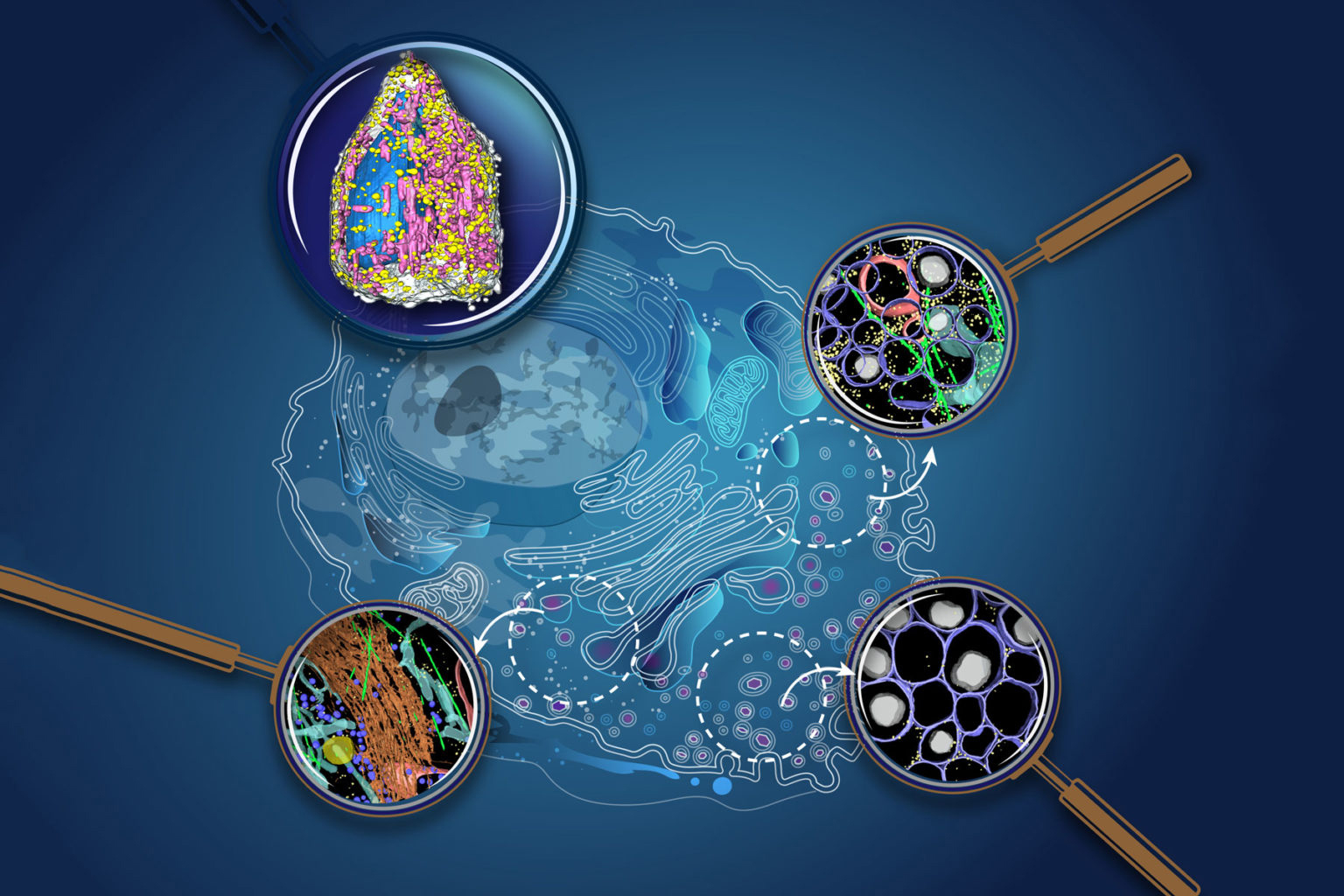
Visualizing the cellular world
Our planet is comprised of continents and islands, each with unique cultures and resources. One area may be well known for growing food, another for manufacturing building materials, and yet despite their differences and distance from one another, the regions are linked by global processes. Living cells are built on a similar concept. For example, one part of the cell produces fuel that powers life, and another part makes the simple building blocks that are then assembled into complex structures inside the cell. To fully understand cells, we need to characterize the structures that make them up, and to identify their contents.
Thanks to advanced imaging technologies, scientists have examined many different components of cells, and some current approaches can even map the structure of these molecules down to each atom. However, getting a glimpse of how all these parts move, change, and interact within a dynamic, living cell has always been a grander challenge.
Merits of soft x-ray tomography
Over the last several decades, the scientific community has unveiled features of cell structure and function using a combination of imaging and biochemical experiments. However, limitations in traditional methods present challenges for quantifying rearrangements and interactions within specific areas inside the cells.
At ALS Beamline 2.1, a team is making waves with a new approach to whole-cell visualization, using the world’s first soft x-ray tomography (SXT) microscope built for biological and biomedical research. SXT is uniquely suited to imaging whole cells without alterations from stains or added tagging molecules, as is the case for fluorescence imaging, and without chemically fixing and sectioning them, which is necessary for traditional electron microscopy.
The ability to analyze intact, near-native-state cells with high throughput allows multiple conditions to be tracked in 3D over time, providing spatially and temporally extended views of integrated cellular responses. Additionally, SXT allows for comparing the molecular densities of organelles before and after drug stimulation, adding new insights into how drugs alter cell function. Using this approach, researchers can directly observe how specific components within a cell respond to stimuli rather than inferring an interaction from biochemical measurements.
Stimulating insulin secretion
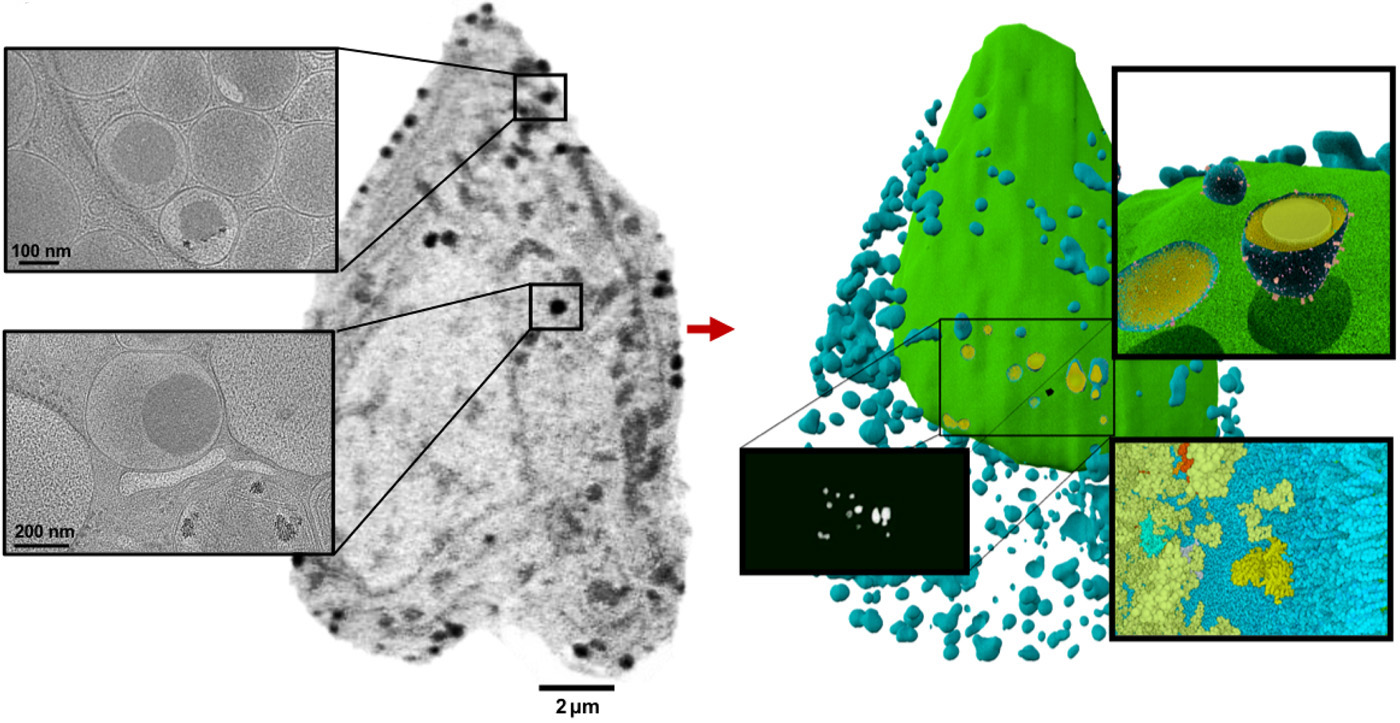
In this study, the team used SXT to reveal never-before-seen details about insulin secretion in pancreatic cells (beta cells) taken from rats—before, during, and after exposure to differing levels of glucose and a common diabetes drug. In rats and other mammals, beta cells respond to rising blood glucose levels by releasing insulin, a hormone that regulates glucose metabolism throughout the body.
The researchers found that stimulating beta cells induced rapid changes in the numbers and molecular densities of insulin vesicles—the membrane “envelopes” in which insulin is stored after production. This was surprising at first, because they expected to see fewer vesicles during secretion, when they are emptied outside the cell. But what they observed was a rapid maturation of existing immature vesicles.
The results not only open the door to the development of improved drugs with fewer side effects, they also represent an important first step toward integrating our knowledge of biology at the molecular and cellular levels, bridging the longstanding gap between structural biology and physiology.
Contact: Carolyn Larabell
Researchers: K.L. White, X. Zhang, J.P. Francis, W. Lin, and K. Tseng (University of Southern California); J. Singla and F. Alber (University of Southern California and University of California Los Angeles); V. Loconte, L. Sun, and A. Li (ShanghaiTech University); J.‑H. Chen, G. McDermott, and C. Larabell (Berkeley Lab and University of California San Francisco); A. Ekman and A. Sali (University of California San Francisco), and R.C. Stevens (University of Southern California and ShanghaiTech University).
Funding: USC Bridge Institute; National Institutes of Health; U.S. Department of Energy (DOE), Office of Biological and Environmental Research; Gordon and Betty Moore Foundation; Chan Zuckerberg Initiative; National Natural Science Foundation of China; and Burroughs Wellcome Fund. Operation of the ALS is supported by DOE, Office of Science, Basic Energy Sciences program.
Publication: K.L. White, J. Singla, V. Loconte, J.‑H. Chen, A. Ekman, L. Sun, X. Zhang, J.P. Francis, A. Li, W. Lin, K. Tseng, G. McDermott, F. Alber, A. Sali, C. Larabell, and R.C. Stevens, “Visualizing subcellular rearrangements in intact cells using soft x-ray tomography,” Sci. Adv. 6, eabc8262 (2020), doi: 10.1126/sciadv.abc8262.
Adapted from the Berkeley Lab news release, “Unique X-Ray Microscope Reveals Dazzling 3D Cell Images.”
ALS SCIENCE HIGHLIGHT #437