by Maritte O’Gallagher
SCIENTIFIC ACHIEVEMENT
Researchers engineered protein-like polymers that replicate complex enzyme functions.
SIGNIFICANCE AND IMPACT
This work, which was verified using X-ray characterization techniques at the Advanced Light Source (ALS), offers a cost-effective, scalable approach that paves the way for functional materials in biomedicine, energy, and manufacturing.
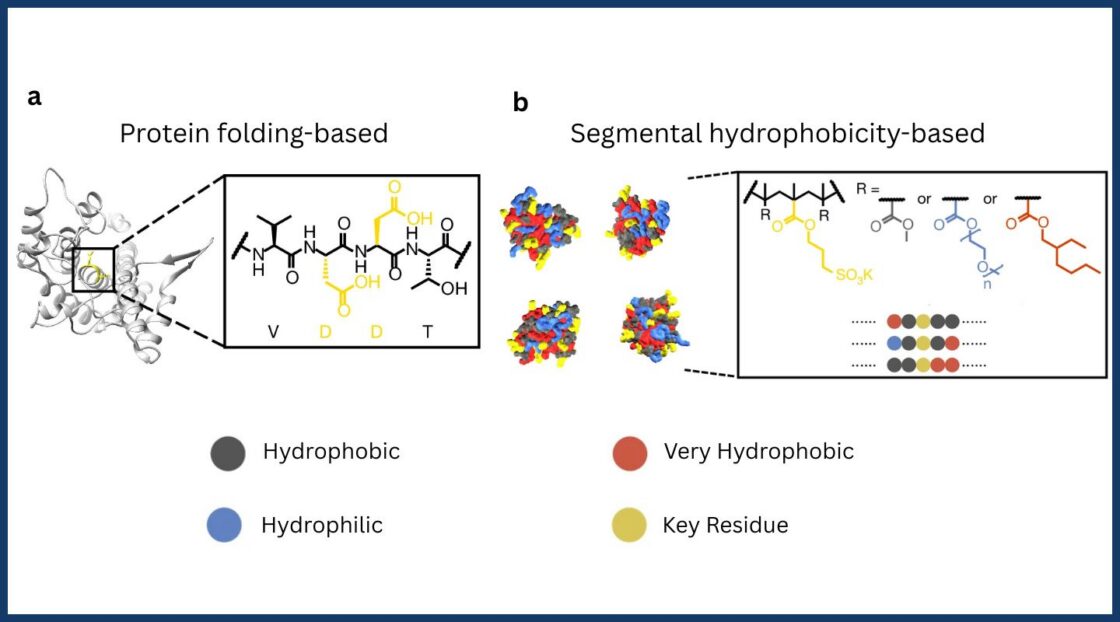
Protein-like functions, without the protein
Many industries already use enzymes, which are specialized protein molecules that accelerate chemical reactions without being consumed. Incorporating these catalytically active molecules into materials could unleash impactful applications biomedicine, energy generation, and chemical synthesis—including masks that eliminate airborne toxicants or environmental filters that degrade pollutants. Their practicality, however, is limited: naturally occurring enzymes tend to be fragile, costly, and unstable.
While these constraints have driven interest in synthetic polymers that mimic enzymatic activity, designing durable protein-like alternatives has been difficult. Natural enzymes rely on rigid secondary structures—local folding patterns along the amino acid chain—that determine whether a target molecule can bind at the active site and trigger a reaction. As a result, past efforts have generally assumed that precise sequence control was necessary to reproduce protein function. This has hindered industrial applications, as specifying the exact order of building blocks in a polymer chain requires costly, high-purity chemical reactions.
In this study, researchers reinterpreted proteins’ sequence-structure-function relationship to engineer polymers with bio-inspired functions and practical adjustments to their molecular chemistry. Using X-ray techniques at the ALS, the team connected how the polymers pack globally with how the local chemical microenvironments near the catalytic region shift upon target binding, a key factor governing function.
Adapting conventional wisdom
The researchers leveraged bioinformatics and machine learning to parse out segments critical to enzyme function and segments that might be less important to precisely control. They then tweaked chunks of the sequence so that the overall polymer would behave like a folded protein. Releasing the need to perfectly control entire sequences, along with using carbon-carbon backbones (instead of the amide backbones of naturally occurring proteins), allowed the researchers to synthesize their protein-like polymers in a simple, efficient, one-pot chemical reaction.
The designed polymers did not form secondary structures but instead adopted varying conformations based on the hydrophobic (water-repelling) properties of segments in the chain. Although the researchers were initially skeptical, this property was ultimately beneficial: the flexible polymers locally adjusted segments to form an impromptu active site, allowing them to bind intended targets and successfully recapitulate the biological effects of natural enzymes.
Still, the researchers had not predesigned the active sites of their polymers, so the molecular interactions underlying this catalytic activity remained unclear. X-ray absorption near-edge structure (XANES) spectroscopy at Beamline 10.3.2, combined with molecular dynamics simulations, probed catalytic site microenvironments, resolving oxidation state, geometry, and hydrophobic interactions. Small-angle X-ray scattering (SAXS) at Beamline 7.3.3, in conjunction with neutron scattering at the High Flux Isotope Reactor and the Spallation Neutron Source at Oak Ridge National Laboratory, uncovered chemical signatures of how polymers altered global folding and rearranged key segments to form functional pseudo-active sites. Additional experiments at the Molecular Foundry characterized polymer structure and composition.
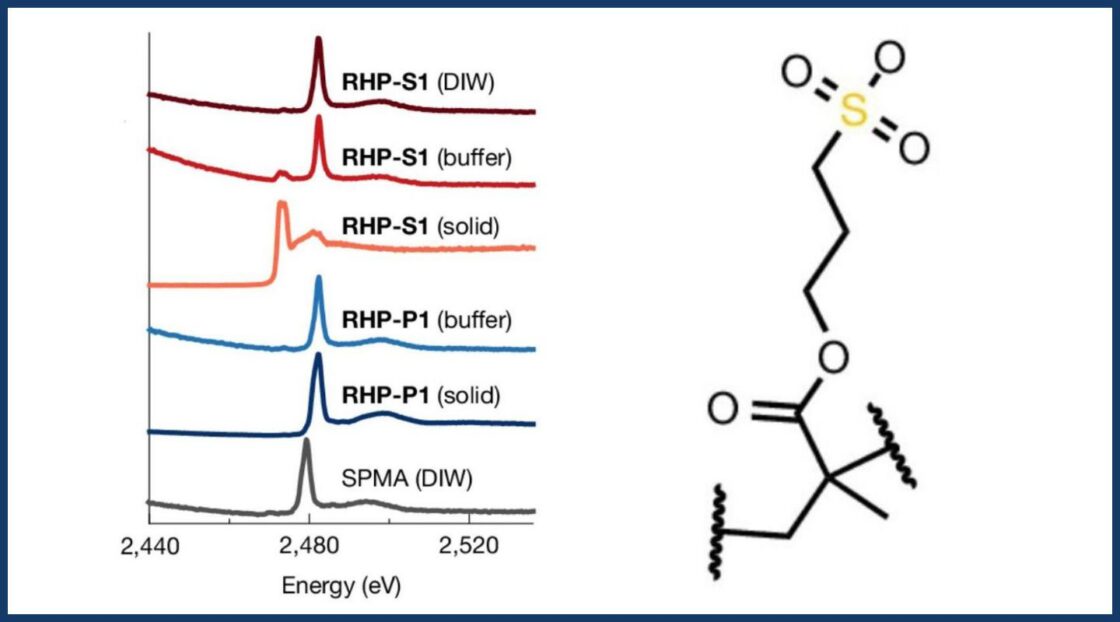
Flexible, scalable design rules for functional materials
Using X-ray and neutron scattering data to validate the mechanisms underlying their approach, the authors demonstrated that it is possible to achieve protein-like function using a statistical approach to sequence control. Built-in flexibility allowed polymers to behave dynamically and compensate for not being perfect replicas of natural enzymes.
Furthermore, the stability of carbon backbones across diverse processing conditions makes these polymers well suited for incorporation into various materials. The research team aims to streamline a design process for applications in energy and biomedicine, like medical-grade antibacterial coatings, drug delivery systems, and custom-engineered tissues. The scalability of their new framework equips the material design community with tools to develop a vast array of protein-like polymers with useful functions.
Contacts: Ting Xu and Jinghua Guo
Researchers: H. Yu, M. Eres, P. Kang, A. Grigoropoulos, D.M. Loh, I. Jayapurna, Z. Ruan, K. Toste, S. Li, H. Huang, and F.D. Toste (University of California, Berkeley); S.L. Hilburg, T. Jin, P. Ganesh, and A. Alexander-Katz (Massachusetts Institute of Technology); Z. Li (University of Illinois Urbana−Champaign); W. Fu and R.D. Britt (University of California, Davis); F. Yang and J. Guo (ALS); Y. Z. (University of Illinois Urbana−Champaign and University of Michigan, Ann Arbor); and T. Xu (University of California, Berkeley and Berkeley Lab).
Funding: Defense Threat Reduction Agency (DTRA); US Department of Defense, Army Research Office; National Science Foundation (NSF); Alfred P. Sloan Foundation, US Department of Energy, Office of Science, Basic Energy Sciences program (DOE BES), Chemical Sciences, Geosciences, and Biosciences Division and Materials Sciences and Engineering Division; DOE Office of Science, Biological Environmental Research program; NSF Graduate Research Fellowship Program; Chyn Duoq Shiah Memorial Fellowship; and the MIT Undergraduate Research Opportunity Program. Operation of the Pines Magnetic Resonance Center’s Core NMR Facility is supported by the National Institutes of Health. Operation of the NIST Center for Neutron Research is supported by the National Institute of Standards and Technology, US Department of Commerce. Operations of the ALS, Molecular Foundry, High Flux Isotope Reactor, and Spallation Neutron Source are supported by the DOE BES.
Publication: H. Yu, M. Eres, S.L. Hilburg, P. Kang, T. Jin, A. Grigoropoulos, Z. Li, D.M. Loh, I. Jayapurna, Z. Ruan, W. Fu, F. Yang, P. Ganesh, K. Toste, S. Li, J. Guo, H. Huang, F.D. Toste, R.D. Britt, Y.Z., A. Alexander-Katz, and T. Xu, “Random heteropolymers as enzyme mimics,” Nature 649, 83 (2026), doi:10.1038/s41586-025-09860-9.
ALS SCIENCE HIGHLIGHT #539