SCIENTIFIC ACHIEVEMENT
Using a multimodal approach that included x-ray scattering at the Advanced Light Source (ALS), researchers determined how components of the SARS-CoV-2 replication mechanism fit together.
SIGNIFICANCE AND IMPACT
A better understanding of how this protein complex works provides insight into potential structural or functional weak spots to exploit for drug development.
A collaboration story

In February 2020, a trio of bio-imaging experts were sitting amiably around a dinner table at a scientific conference in Washington, D.C., when the conversation shifted to what was then a worrying viral epidemic in China. Without foreseeing the global disaster to come, they wondered aloud how they might contribute.
Nearly a year and a half later, Greg Hura of Berkeley Lab, Andrzej Joachimiak of Argonne National Laboratory, and Hugh M. O’Neill of Oak Ridge National Laboratory, plus their many collaborators, have published a comprehensive study that—alongside other recent, complementary studies of coronavirus proteins and genetics—represents the first step toward developing treatments for that viral infection, now seared into the global consciousness as COVID-19.
The replication transcription complex
The protein-based machine responsible for RNA replication and translation in coronaviruses—and many other viruses—is called the replication transcription complex (RTC). To successfully duplicate viral RNA for new virus particles and produce the new particles’ many proteins, the RTC must: distinguish between viral and host RNA, recognize and pair RNA bases instead of highly similar DNA bases that are also abundant in human cells, convert their RNA into mRNA (to dupe human ribosomes into translating viral proteins), interface with error-checking molecules, and transcribe specific sections of viral RNA to amplify certain proteins over others depending on need—all while trying to evade the host immune system.
Multitasking proteins like the RTC aren’t static or rigid; they’re flexible and have associated molecules, called nonstructural proteins, that change form depending on the task at hand. Each arrangement exposes different parts of the RTC surface, which can be examined to find places where potential drug molecules could bind and inhibit the entire machine.
From structure to function
In this multimodal study, the researchers used a combination of x-ray crystallography at the Advanced Photon Source, small-angle neutron scattering at Oak Ridge’s High Flux Isotope Reactor and Spallation Neutron Source, and small-angle x-ray scattering (SAXS) at ALS Beamline 12.3.1. The latter technique is particularly useful for probing the transient and plastic properties of assemblies in solution. The structures and interactions of three nonstructural proteins that make up RTC (Nsp7, Nsp8, and Nsp12) were determined in various combinations, with and without RNA.
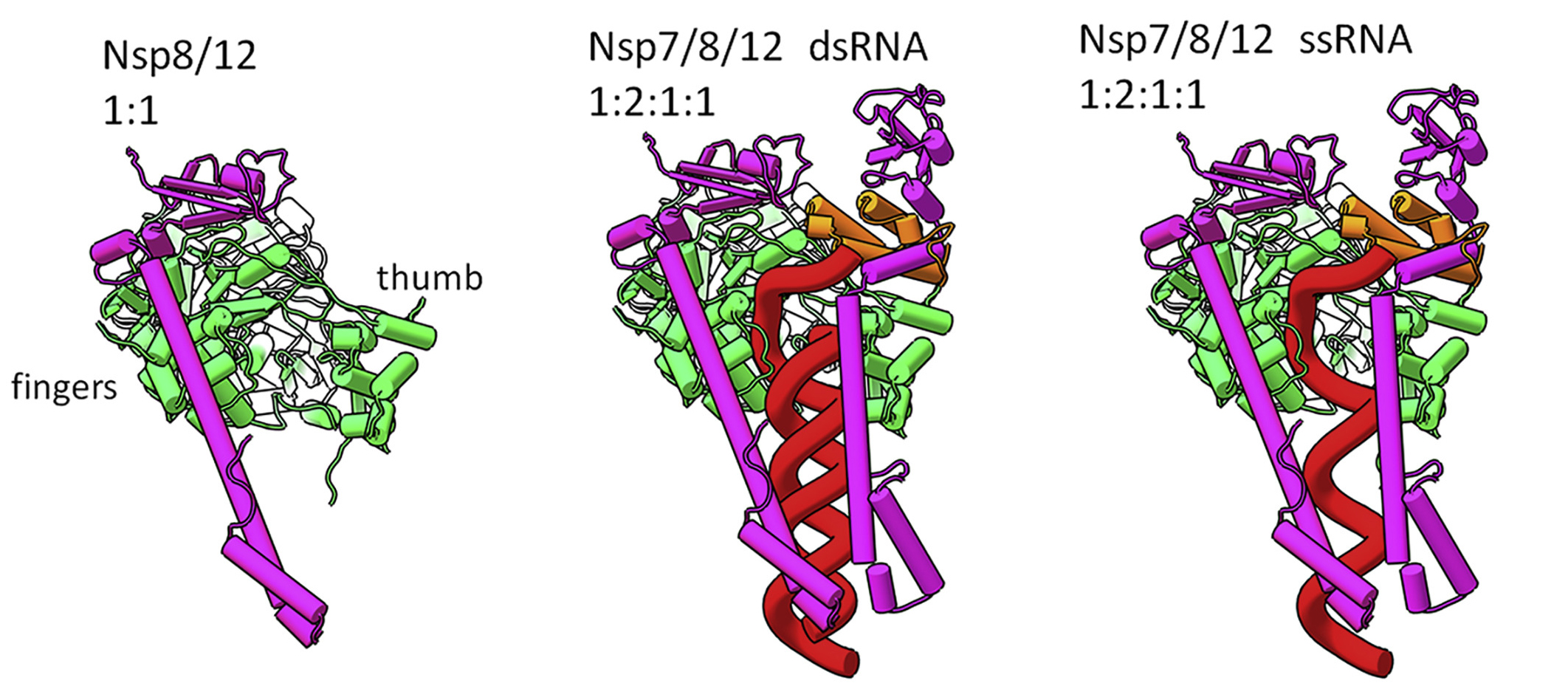
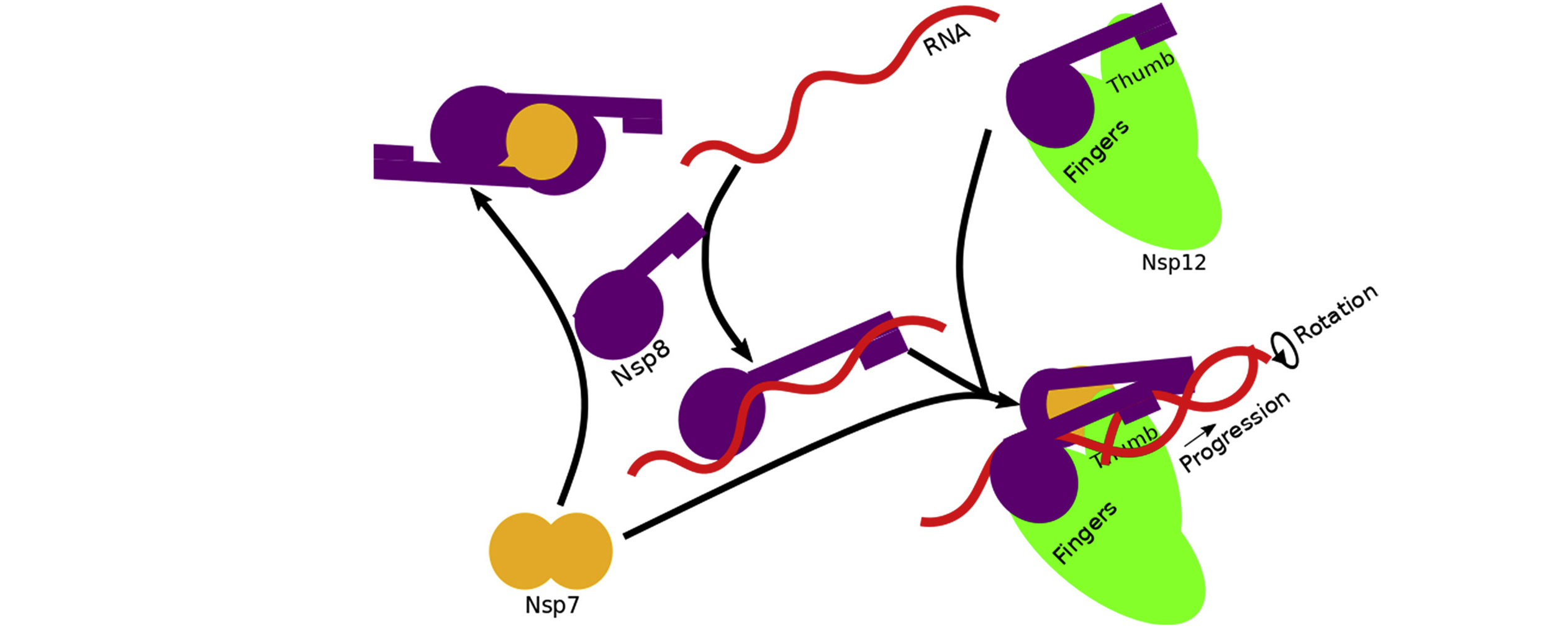
The data revealed that assembly of the RTC is incredibly precise. The nonstructural proteins can’t move into place randomly; they must follow a specific order. For example, before RNA transcription can occur, Nsp7, Nsp8, and RNA must combine with a stabilized complex of Nsp8/Nsp12. Based on the architecture of the resulting complex, the researchers proposed a mechanism in which RNA rotation and progression pulls one of the two Nsp8 subunits off of its spot on Nsp12 (the “fingers”), leaving a vacancy to be filled by the remaining Nsp8 (bound to the “thumb”).
Due to the similarity of RTC proteins across viral strains, the researchers believe that any drugs developed to block its activity could work for multiple viral infections in addition to all COVID-19 variants. However, a greater understanding of the RTC machinery is still needed to provide insight into how current drugs work and to suggest other ways of interfering with its function.
Contact: Greg Hura
Researchers: M. Wilamowski (University of Chicago); M. Hammel, J. Bierma, A.H. Sarker, and S.E. Tsutakawa (Berkeley Lab); W. Leite, Q. Zhang, K. Weiss, Y. Fan, S.V. Pingali, and H.M. O’Neill (Oak Ridge National Laboratory); Y. Kim, R. Jedrzejczak, and A. Joachimiak (University of Chicago and Argonne National Laboratory); D.J. Rosenberg (Berkeley Lab and University of California, Berkeley); J. Wower (Auburn University); and G.L. Hura (Berkeley Lab and University of California, Santa Cruz).
Funding: U.S. Department of Energy (DOE) National Virtual Biotechnology Laboratory, a consortium of DOE National Laboratories with core capabilities relevant to the threats posed by COVID-19, and funded under the Coronavirus Aid, Relief, and Economic Security (CARES) Act; DOE Biological and Environmental Research Integrated Diffraction Analysis Technologies program; National Institutes of Health; United States–Israel Binational Science Foundation; U.S. Department of Agriculture; DOE Office of Science, Basic Energy Sciences (BES) program. Operation of the ALS is supported by DOE BES.
Publication: M. Wilamowski, M. Hammel, W. Leite, Q. Zhang, Y. Kim, K. Weiss, R. Jedrzejczak, D.J. Rosenberg, Y. Fan, J. Wower, J. Bierma, A.H. Sarker, S.E. Tsutakawa, S.V. Pingali, H.M. O’Neill, A. Joachimiak, and G.L. Hura, “Transient complexes of the Nsp7, Nsp8 and Nsp12 in SARS-CoV-2 replication transcription complex,” Biophys. J. 120, 3152 (2021), doi: 10.1016/j.bpj.2021.06.006.
Adapted from the Berkeley Lab news release: “Deconstructing the Infectious Machinery of the SARS-CoV-2 Virus.”
ALS SCIENCE HIGHLIGHT #449