In the struggle to keep up with microbes whose rapid mutations outpace our ability to produce vaccines, the human race has a powerful ally: computers. Researchers have now figured out a way to use computational protein design to generate small, stable proteins that accurately mimic key viral structures; these can then be used in vaccines to induce potent neutralizing antibodies. The results were validated in part using protein structures obtained at several of the Berkeley Center for Structural Biology protein crystallography beamlines at the ALS. The results have yielded promising leads for the development of vaccines to protect infants, young children, and the elderly against respiratory illness. More generally, the results provide proof of principle for this approach, which can apply to a variety of other vaccine targets, such as human immunodeficiency virus and influenza.
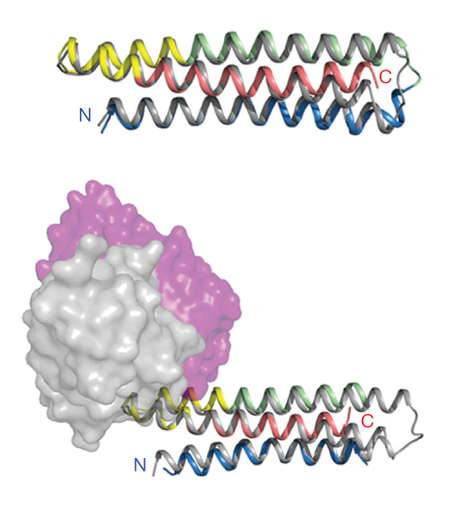
The researchers in this study used the fact that there are patches on pathogens that antibodies recognize: the “fingerprints,” or in biological terms, the “epitopes.” Using this new method called “epitope-focused vaccine design,” the scientists first isolated antibodies from patients infected with RSV (respiratory syncytial virus), a virus resistant to current strategies for designing vaccines and that is responsible for an estimated annual six to seven percent of deaths of children under age one. Then, working backward from those antibodies, they determined the epitope that the antibodies recognize. It was important in this case to use a class of antibodies called “broadly neutralizing,” which means that they recognize a key epitope on the virus—a region that the antibodies recognize no matter what mutations might occur in other regions of the pathogen. Often, viruses hide these key epitopes from detection by antibodies, which is part of the reason it is difficult to design vaccines against them.
The researchers then took that epitope, the unchanging fingerprint of the virus, and computationally modeled it into various protein scaffolds, designing a virtual virus. Computationally designing a protein is no easy task: for many years researchers have tried to design proteins de novo, but when the designed proteins are grown in cells, they often lack functional ability. The scientists in this study used a new approach in that they allowed much greater flexibility in their scaffold protein structure, allowing the computationally modeled protein to “virtually” fold in many different ways around the epitope. They then used energy-minimizing algorithms to select the most stable structures.
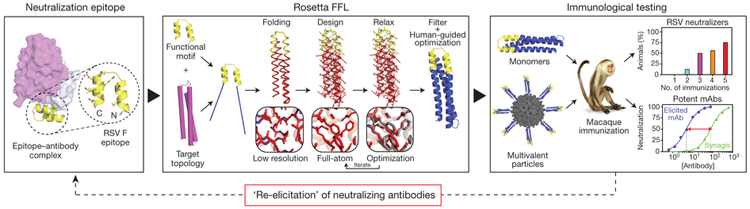
This computationally intensive approach paid off. When the resulting structures were expressed in and purified from E. coli cells, they behaved like normal proteins with good functional activity, and even better, they showed excellent affinity for the human antibodies for which they were designed. Finally, testing in rhesus macaques showed that the animals produced antibodies in response to the designed viruses.
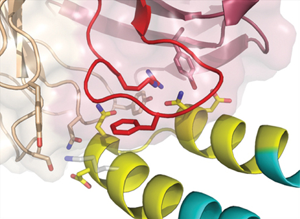
But did these computationally designed proteins really grow into the structures that the researchers had designed? This is where protein crystallography comes in; crystallography is the foremost technique to look at protein structure on an atomic level. The designed proteins were crystallized and the structures determined at ALS Beamlines 5.0.1, 5.0.2, and 8.2.2, which not only gave the final proof that the proteins were exactly what the researchers had designed, but also gave deeper insight into why and how they bound to the antibodies.
This new method—computationally designing a “wanted poster” for the immune system—promises to open up a whole new way of making vaccines. Designed with sophisticated computational routines, grown in a lab, and then structurally determined using crystallography, the designed pathogen can mimic a real pathogen well enough that the body is tricked into producing protective antibodies, so that when and if the real pathogen shows up, the immune system is ready. The days of using dead viruses against themselves may soon be over.
Contact: William Schief
Research conducted by: G. Baneyx, V. Vittal, E. Stevens, A. Schroeter, R.E. Klevit, and D. Baker (Univ. of Washington); B.E. Correia (U. of Washington; Univ. Nova de Lisboa, Portugal; and Scripps Research Institute); J.T. Bates and J.E. Crowe (Vanderbilt University Medical Center); R.J. Loomis, M.J. Connell, and P.R. Johnson (Children’s Hospital of Philadelphia Research Institute); C. Carrico, P. Rupert, C. Correnti, M.A. Holmes, and R.K. Strong (Fred Hutchinson Cancer Research Center); J.G. Jardine, O. Kalyuzhniy, S. MacPherson, A.M. Serra, Y. Adachi, and W.R. Schief (U. of Washington and Scripps Research Institute); Y. Li and R.T. Wyatt (Scripps Research Institute); M. Chen and B.S. Graham (National Institutes of Health).
Research funding: Foundation for Science and Technology, Portugal; National Institutes of Health; Children’s Hospital of Philadelphia; Bill and Melinda Gates Foundation; International AIDS Vaccine Initiative; and March of Dimes. Operation of the ALS is supported by the U.S. Department of Energy, Office of Basic Energy Sciences.
Publication about this research: B.E. Correia, J.T. Bates, R.J. Loomis, G. Baneyx, C. Carrico, J.G. Jardine, P. Rupert, C. Correnti, O. Kalyuzhniy, V. Vittal, M.J. Connell, E. Stevens, A. Schroeter, M. Chen, S. MacPherson, A.M. Serra, Y. Adachi, M.A. Holmes, Y. Li, R.E. Klevit, B.S. Graham, R.T. Wyatt, D. Baker, R.K. Strong, J.E. Crowe, P.R. Johnson, and W.R. Schief, “Proof of principle for epitope-focused vaccine design,” Nature 507, 201 (2014). doi: 10.1038/nature12966
ALS SCIENCE HIGHLIGHT #297