by Lori Tamura
 In 2011, Jennifer Doudna and Emmanuelle Charpentier met at a scientific conference in Puerto Rico. Wandering through the streets of old San Juan, they talked about how interesting it would be to work on a rather esoteric subject: how a protein, known as CRISPR-Cas9, protects bacteria from viral infections. As Doudna described in an interview posted by UC Berkeley, it was “a collaboration between two women who set out, not to win the Nobel Prize, but to do really fundamental science that we thought was exciting and important and interesting.”
In 2011, Jennifer Doudna and Emmanuelle Charpentier met at a scientific conference in Puerto Rico. Wandering through the streets of old San Juan, they talked about how interesting it would be to work on a rather esoteric subject: how a protein, known as CRISPR-Cas9, protects bacteria from viral infections. As Doudna described in an interview posted by UC Berkeley, it was “a collaboration between two women who set out, not to win the Nobel Prize, but to do really fundamental science that we thought was exciting and important and interesting.”
That meeting in Puerto Rico marked the beginning of a remarkable journey for Doudna, professor of molecular and cell biology and chemistry and founder of the Innovative Genomics Institute at UC Berkeley, and Charpentier, founding, scientific and managing director of the Max Planck Unit for the Science of Pathogens, Berlin. Just nine years after first crossing paths, they have reached the pinnacle of scientific achievement: the Nobel Prize for Chemistry in 2020. Their development of extremely precise, programmable “genetic scissors” for editing DNA revolutionized biomedicine and life sciences and offers promising new approaches to solving energy and environmental challenges.
It is a momentous achievement, supported by the contributions of numerous individuals and institutions, including the Advanced Light Source (ALS) synchrotron user facility at Lawrence Berkeley National Laboratory (Berkeley Lab). “Jennifer’s CRISPR-Cas9 crystallographic work has been performed almost exclusively at the ALS structural biology beamlines,” said Paul Adams, director of Berkeley Lab’s Molecular Biophysics and Integrated Bioimaging (MBIB) Division, where Doudna is a faculty scientist. “I am delighted that her outstanding research has received the ultimate scientific recognition.”
The seeds for this success were planted at the ALS in the 1990s, when forward thinkers began a protein crystallography program—based on hard x-rays—at a facility optimized for soft x-rays. “Now the wisdom of those efforts is even more broadly recognized,” said ALS Director Steve Kevan. “We extend our congratulations to Jennifer for her monumental achievement and to the ALS and Berkeley Lab staff who contributed to making this possible.”
Understanding how the molecules work
 The “eureka moment” for Doudna and her then-postdoc, Martin Jinek, came when they realized that they could control where CRISPR-Cas9 cuts DNA by changing the RNA snippet that guides it to a particular DNA sequence. “What made CRISPR so clearly special is that it’s easily deployable,” said Doudna at a press conference on the morning of the Nobel announcement. “Understanding the molecular basis of how it works as a bacterial defense system immediately made it clear this would be a very useful tool in other kinds of cells.”
The “eureka moment” for Doudna and her then-postdoc, Martin Jinek, came when they realized that they could control where CRISPR-Cas9 cuts DNA by changing the RNA snippet that guides it to a particular DNA sequence. “What made CRISPR so clearly special is that it’s easily deployable,” said Doudna at a press conference on the morning of the Nobel announcement. “Understanding the molecular basis of how it works as a bacterial defense system immediately made it clear this would be a very useful tool in other kinds of cells.”
Hard x-ray crystallography is the technique of choice for scientists who want to gain an atomic-level understanding of molecular structure. In the dozen years leading up to the aforementioned eureka moment, Doudna had become well acquainted with the ALS, which was ramping up its crystallography capabilities at the time. In 2000, while still a professor at Yale, she published her first ALS-linked paper, utilizing the wiggler-based Beamline 5.0.2. In 2002, she transferred to UC Berkeley—just a five-minute drive from the ALS—and quickly got involved in the ALS community, joining the Scientific Advisory Committee and serving as chair of the Users’ Executive Committee in 2003.
Over the ensuing years, Doudna published 35 papers using ALS crystallography beamlines, primarily the three Sector 8 superbend beamlines: the joint UC Berkeley/UCSF Beamline 8.3.1 and the Howard Hughes Medical Institute (HHMI) Beamlines 8.2.1 and 8.2.2. Doudna was particularly interested in understanding the molecular basis for genetic control—specifically, the ways in which RNA molecules affect the flow of genetic information in cells. The Nobel Committee cited two of Doudna’s ALS-supported papers (Jinek 2014 and Jiang 2015) in its CRISPR-Cas9 scientific background document.
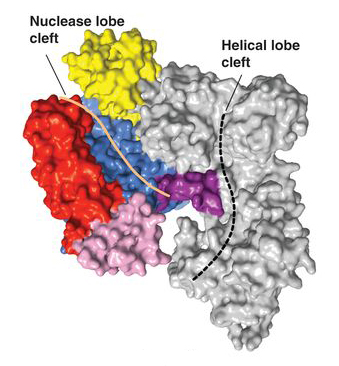
In the first paper, Cas9 structures from two types of bacteria were solved, revealing the common structural core shared by all Cas9 enzymes: an open, two-lobed architecture with clefts for binding nucleic acids. Moreover, the work showed that structural changes are required to trigger DNA recognition and cleavage. (See also the ALS science highlight, “Intriguing DNA Editor Has a Structural Trigger.”) In the second paper, the researchers reported that, when Cas9 binds to a DNA-targeting guide RNA, it adopts a new conformation, distinct from both its unbound and DNA-bound states. In addition, the guide RNA assumes a wider, more compact form that prepares it to recognize the DNA target sequence. Other ALS highlights written about Doudna’s CRISPR work include “Foreign DNA Capture during CRISPR–Cas Adaptive Immunity” and “The CRISPR Target-Recognition Mechanism.”
A detailed structural understanding of how Cas9 recognizes its DNA targets and mediates cleavage has been important for efforts to engineer improved versions of the system. In an ALS feature story about a CRISPR-based biotech company founded in 2011 (“Caribou Biosciences Has Roots at the ALS”), Doudna is quoted saying, “Having the ALS available to us to be able to look at the molecular structures of these proteins has been absolutely critical for Caribou. Without a molecular understanding it would be impossible to do what we do.”
The roots of ALS structural biology
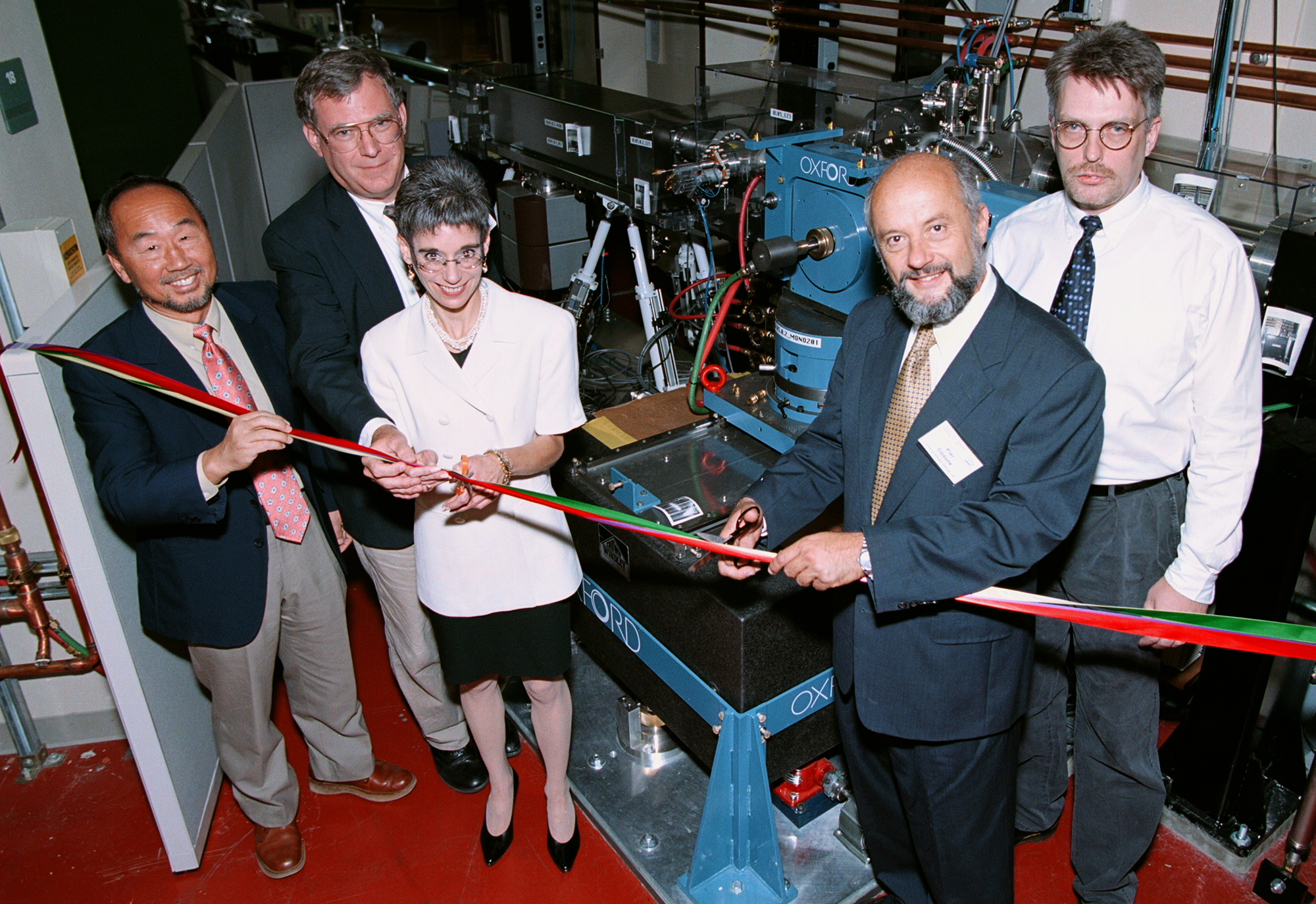

The first structural biology beamline at the ALS, the wiggler-based Beamline 5.0.2, came online in 1997, designed by ALS scientist Howard Padmore and biophysicist Thomas Earnest. The beamline was a spectacular success, and protein crystallographers were soon clamoring for more beamtime. “It was immediately obvious that we needed more capacity and perhaps even higher brightness,” said Padmore. Demonstration experiments—performed by ALS scientist Alastair MacDowell and MBIB biophysicists Bob Glaeser and James Holton—showed that protein crystallography was possible on bend-magnet beamlines, even at a low-energy facility such as the ALS.
By the late 1990s, Chuck Shank (then Berkeley Lab director) and Graham Fleming (director of the Physical Biosciences Division), with support from UC Berkeley leadership, were eager to rapidly expand the structural biology program in Berkeley, following the outstanding success of the wiggler-based beamline for protein crystallography. Since expansion with other wigglers was not possible, high-performance bend-magnet sources offered a highly innovative solution. “When the superbend idea was presented, we jumped on it,” said Shank.
Superbends: high risk, high reward
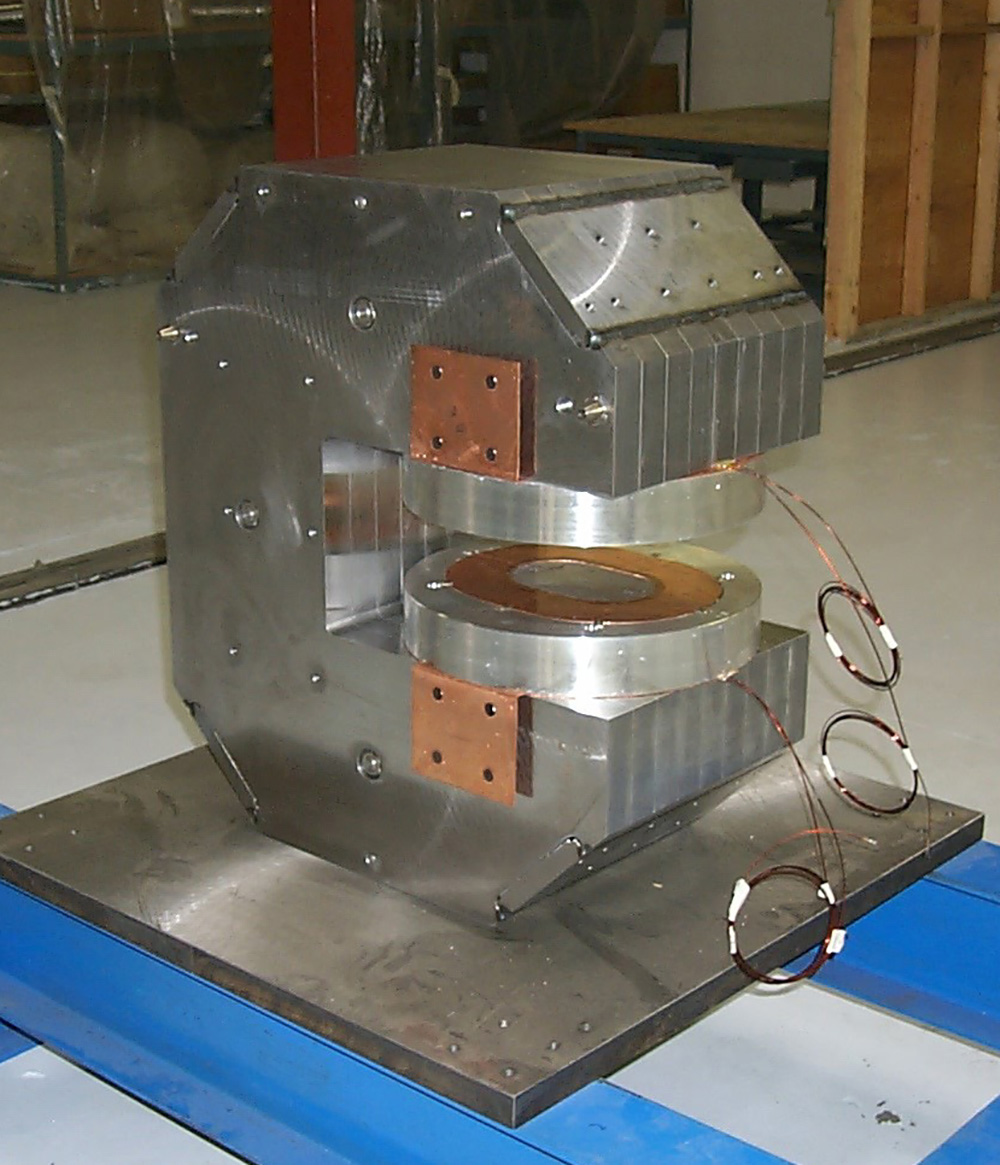

Superconducting bend magnets (superbends)—high-field magnets that enable the generation of harder x-rays in a low-energy storage ring such as the ALS—were first discussed in the early 1990s by Alan Jackson, who led the ALS Accelerator Physics Group (and who, sadly, recently passed away). In that era, the use of superbends in synchrotron light sources was revolutionary and risky. Unlike straight-section insertion devices such as wigglers and undulators, superbends cannot simply be switched off in case of malfunction; they must work to steer electrons around the storage ring so that other beamlines can continue to operate.
Berkeley Lab accelerator physicist Dave Robin, now director of the ALS Upgrade Project, conducted preliminary modeling studies to ensure that superbends would work. Clyde Taylor, a pioneer of superconducting magnet technology, led a Laboratory Directed Research and Development (LDRD) project to design and build a superbend prototype. Ultimately, with Robin’s leadership, the superbends were successfully brought online in 2001.
The beamlines bear fruit
On the beamline side, user Tom Alber (from UCB) spearheaded efforts to secure public and private funding for the ALS’s first superbend beamline (8.3.1) for protein crystallography, including support from UCB and (with user David Agard) UCSF. The beamline and endstation designs, based on innovations developed by Padmore and Holton, were instrumental in convincing representatives of HHMI, brought in by Fleming, to fund two more superbend beamlines (8.2.1 and 8.2.2).
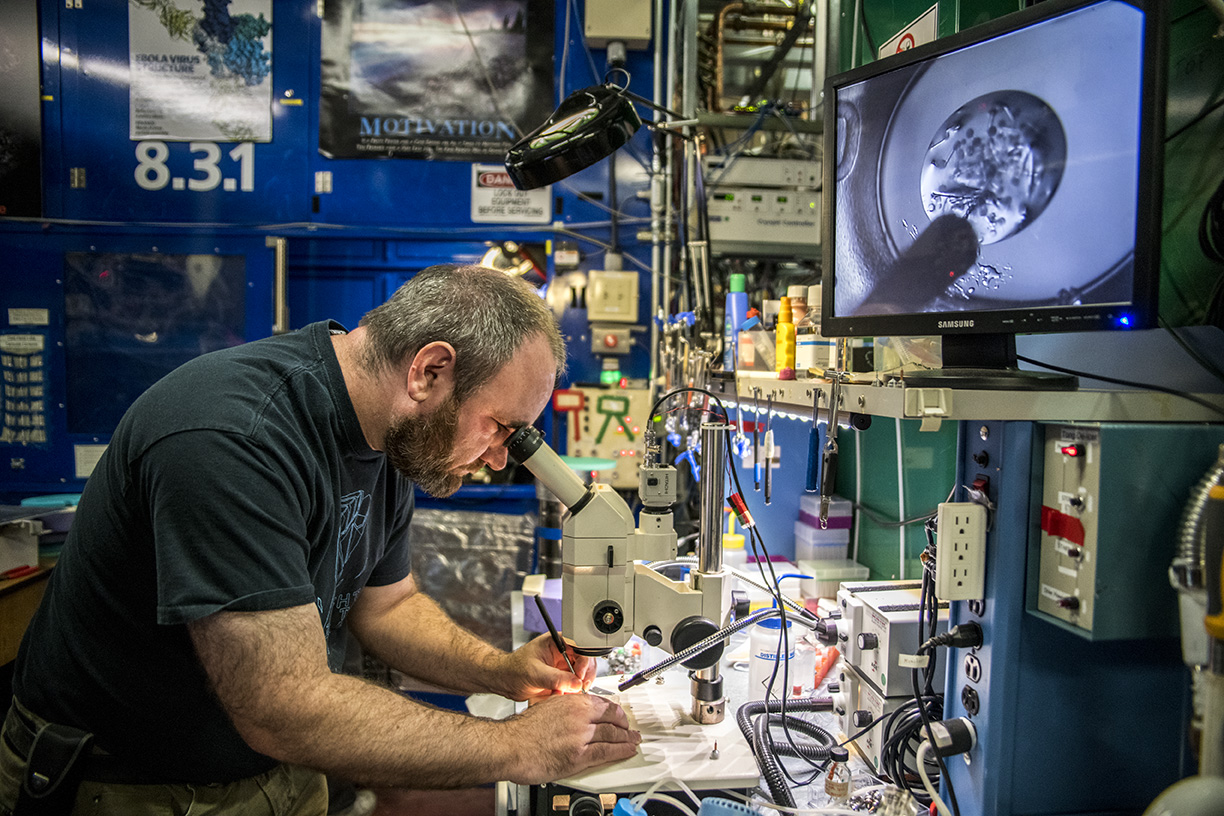

As a UC Berkeley faculty member and HHMI investigator, Doudna had ready access to these facilities. Soon, she began publishing a steady stream of papers from these beamlines in high-profile journals, covering numerous topics involving CRISPR-Cas proteins, of course, but also hepatitis delta virus (HDV), HIV-1, and cholera bacteria.
“Jennifer knows how to recruit really smart people and get them working on really hard, but also really important, scientific problems,” said Holton, now the 8.3.1 beamline scientist. “CRISPR was very challenging to work with in the beginning, but now that Jennifer and others like her have figured out how to use it, it’s a powerful tool for all.”
The ALS protein crystallography program today also includes wiggler side-branch Beamlines 5.0.1 and 5.0.3 and superbend Beamlines 4.2.2 and 12.3.1, with the first hard x-ray undulator Beamline 2.0.1 (“Gemini”) being commissioned this year. The beamlines are extremely productive, collectively depositing over 8000 structures into the Protein Data Bank and shedding light on diseases having broad societal impact, including COVID-19, tuberculosis, cancer, influenza, Zika, and Ebola.
Much of the work is fundamental in nature and not so narrowly associated with a particular disease or condition. Such studies ask questions about how proteins pack and unpack DNA in cells in response to stressful envirnoments, what forces stabilize membrane proteins, and even how to design novel proteins from scratch.
“Solutions to problems come from unexpected directions,” said Doudna in the UC Berkeley interview. “I always encourage students to pursue their passions, because we don’t know where the next big discoveries and technologies are going to come from. Who knew that a bacterial immune system would in fact emerge as a world-changing technology for gene editing? But here we are.”